
AI in health and medicine is a game-changer, revolutionizing diagnostics, treatment, and healthcare management with a focus on predictive analytics and personalized medicine, marking a transformative impact at the forefront of medical innovation.
a. Nucleus Detection and Segmentation
Related Publications
b Nucleus Classification
Related Publications
c. Tissue Phenotyping
Related Publications
● Knowledge Distillation in Histology Landscape by Multi-Layer Features Supervision, IEEE Journal of Biomedical and Health Informatics, 2023
● Multiplex Cellular Communities in Multi-Gigapixel Colorectal Cancer Histology Images for Tissue Phenotyping, IEEE Transactions on Image Processing, 2020
● Cellular Community Detection for Tissue Phenotyping In Colorectal Cancer Histology Images, Medical Image Analysis, 2020
● CPLIP: Zero-Shot Learning for Histopathology with Comprehensive Vision-Language Alignment, CVPR, 2024
d. Skin Cancer Segmentation
This work introduces an innovative and lightweight method for foot ulcer segmentation, setting it apart from established techniques in the field. The proposed model integrates residual connections, channel attention, and spatial attention within convolution blocks. Notably, our approach diverges from widely used backbone architectures, pre-training, and transfer learning strategies. The training methodology employed is a straightforward patch-based approach, enriched with test time augmentations and majority voting. This simplicity in training underscores the effectiveness of our lightweight solution, providing a compelling alternative to the more intricate architectures commonly utilized in foot ulcer segmentation tasks.

Related Publications
e. Prostate Cancer Detection
Related Publications
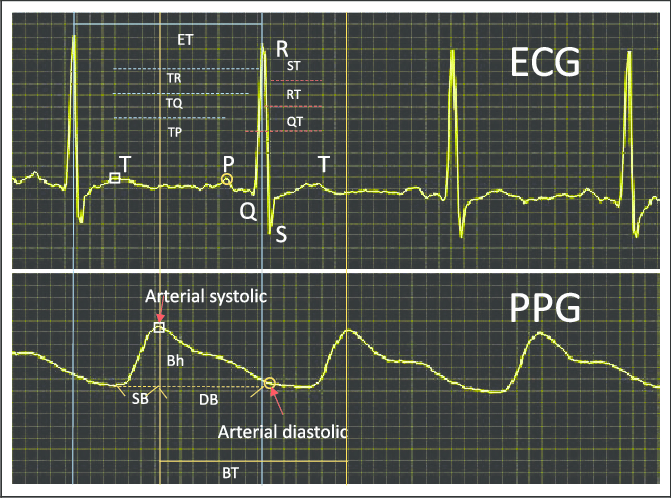
f. ECG Estimation Using Low-Cost Portable PPG Sensors
g. Emotion Detection Using ECG Signal
Related Publications
h. Classification of Benign and Malignant Masses in Mammograms
Related Publications
i. Unsupervised WSI Classification
Related Publications
● CPLIP: Zero-Shot Learning for Histopathology with Comprehensive Vision-Language Alignment, CVPR, 2024
